Image#
The Image class, representing one medical image,
stores a 4D tensor, whose voxels encode, e.g., signal intensity or segmentation
labels, and the corresponding affine transform,
typically a rigid (Euclidean) transform, to convert
voxel indices to world coordinates in mm.
Arbitrary fields such as acquisition parameters may also be stored.
Subclasses are used to indicate specific types of images,
such as ScalarImage and LabelMap,
which are used to store, e.g., CT scans and segmentations, respectively.
An instance of Image can be created using a filepath,
a PyTorch tensor, or a NumPy array.
This class uses lazy loading, i.e., the data is not loaded from disk at
instantiation time.
Instead, the data is only loaded when needed for an operation
(e.g., if a transform is applied to the image).
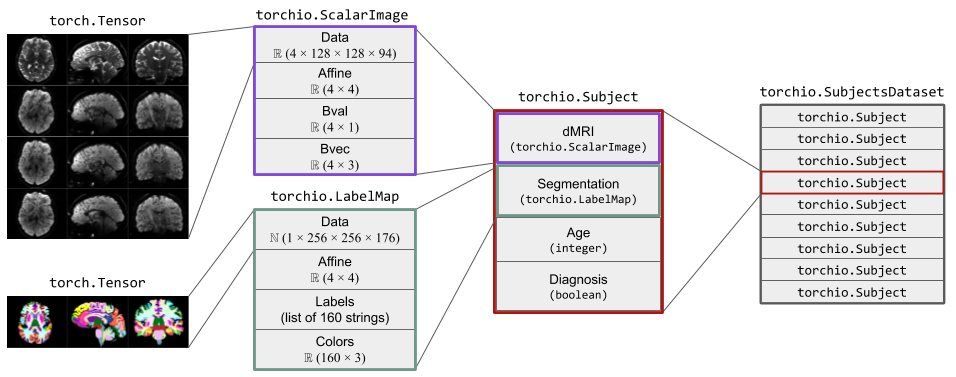
The figure below shows two instances of Image.
The instance of ScalarImage contains a 4D tensor representing a
diffusion MRI, which contains four 3D volumes (one per gradient direction),
and the associated affine matrix.
Additionally, it stores the strength and direction for each of the four
gradients.
The instance of LabelMap contains a brain parcellation of the same
subject, the associated affine matrix, and the name and color of each brain
structure.
- class torchio.ScalarImage(*args, **kwargs)[source]#
Bases:
ImageImage whose pixel values represent scalars.
Example
>>> import torch >>> import torchio as tio >>> # Loading from a file >>> t1_image = tio.ScalarImage('t1.nii.gz') >>> dmri = tio.ScalarImage(tensor=torch.rand(32, 128, 128, 88)) >>> image = tio.ScalarImage('safe_image.nrrd', check_nans=False) >>> data, affine = image.data, image.affine >>> affine.shape (4, 4) >>> image.data is image[tio.DATA] True >>> image.data is image.tensor True >>> type(image.data) torch.Tensor
See
Imagefor more information.
- class torchio.LabelMap(*args, **kwargs)[source]#
Bases:
ImageImage whose pixel values represent categorical labels.
Example
>>> import torch >>> import torchio as tio >>> labels = tio.LabelMap(tensor=torch.rand(1, 128, 128, 68) > 0.5) >>> labels = tio.LabelMap('t1_seg.nii.gz') # loading from a file >>> tpm = tio.LabelMap( # loading from files ... 'gray_matter.nii.gz', ... 'white_matter.nii.gz', ... 'csf.nii.gz', ... )
Intensity transforms are not applied to these images.
Nearest neighbor interpolation is always used to resample label maps, independently of the specified interpolation type in the transform instantiation.
See
Imagefor more information.
- class torchio.Image(path: str | ~pathlib.Path | ~typing.Sequence[str | ~pathlib.Path] | None = None, type: str | None = None, tensor: ~torch.Tensor | ~numpy.ndarray | None = None, affine: ~torch.Tensor | ~numpy.ndarray | None = None, check_nans: bool = False, reader: ~typing.Callable = <function read_image>, **kwargs: ~typing.Dict[str, ~typing.Any])[source]#
Bases:
dictTorchIO image.
For information about medical image orientation, check out NiBabel docs, the 3D Slicer wiki, Graham Wideman’s website, FSL docs or SimpleITK docs.
- Parameters:
path – Path to a file or sequence of paths to files that can be read by
SimpleITKornibabel, or to a directory containing DICOM files. Iftensoris given, the data inpathwill not be read. If a sequence of paths is given, data will be concatenated on the channel dimension so spatial dimensions must match.type – Type of image, such as
torchio.INTENSITYortorchio.LABEL. This will be used by the transforms to decide whether to apply an operation, or which interpolation to use when resampling. For example, preprocessing and augmentation intensity transforms will only be applied to images with typetorchio.INTENSITY. Spatial transforms will be applied to all types, and nearest neighbor interpolation is always used to resample images with typetorchio.LABEL. The typetorchio.SAMPLING_MAPmay be used with instances ofWeightedSampler.tensor – If
pathis not given,tensormust be a 4Dtorch.Tensoror NumPy array with dimensions \((C, W, H, D)\).affine – \(4 \times 4\) matrix to convert voxel coordinates to world coordinates. If
None, an identity matrix will be used. See the NiBabel docs on coordinates for more information.check_nans – If
True, issues a warning if NaNs are found in the image. IfFalse, images will not be checked for the presence of NaNs.reader – Callable object that takes a path and returns a 4D tensor and a 2D, \(4 \times 4\) affine matrix. This can be used if your data is saved in a custom format, such as
.npy(see example below). If the affine matrix isNone, an identity matrix will be used.**kwargs – Items that will be added to the image dictionary, e.g. acquisition parameters.
TorchIO images are lazy loaders, i.e. the data is only loaded from disk when needed.
Example
>>> import torchio as tio >>> import numpy as np >>> image = tio.ScalarImage('t1.nii.gz') # subclass of Image >>> image # not loaded yet ScalarImage(path: t1.nii.gz; type: intensity) >>> times_two = 2 * image.data # data is loaded and cached here >>> image ScalarImage(shape: (1, 256, 256, 176); spacing: (1.00, 1.00, 1.00); orientation: PIR+; memory: 44.0 MiB; type: intensity) >>> image.save('doubled_image.nii.gz') >>> def numpy_reader(path): ... data = np.load(path).as_type(np.float32) ... affine = np.eye(4) ... return data, affine >>> image = tio.ScalarImage('t1.npy', reader=numpy_reader)
- as_pil(transpose=True)[source]#
Get the image as an instance of
PIL.Image.Note
Values will be clamped to 0-255 and cast to uint8.
Note
To use this method, Pillow needs to be installed:
pip install Pillow.
- axis_name_to_index(axis: str) int[source]#
Convert an axis name to an axis index.
- Parameters:
axis – Possible inputs are
'Left','Right','Anterior','Posterior','Inferior','Superior'. Lower-case versions and first letters are also valid, as only the first letter will be used.
Note
If you are working with animals, you should probably use
'Superior','Inferior','Anterior'and'Posterior'for'Dorsal','Ventral','Rostral'and'Caudal', respectively.Note
If your images are 2D, you can use
'Top','Bottom','Left'and'Right'.
- property data: Tensor#
Tensor data (same as
Image.tensor).
- static flip_axis(axis: str) str[source]#
Return the opposite axis label. For example,
'L'->'R'.- Parameters:
axis – Axis label, such as
'L'or'left'.
- classmethod from_sitk(sitk_image)[source]#
Instantiate a new TorchIO image from a
sitk.Image.Example
>>> import torchio as tio >>> import SimpleITK as sitk >>> sitk_image = sitk.Image(20, 30, 40, sitk.sitkUInt16) >>> tio.LabelMap.from_sitk(sitk_image) LabelMap(shape: (1, 20, 30, 40); spacing: (1.00, 1.00, 1.00); orientation: LPS+; memory: 93.8 KiB; dtype: torch.IntTensor) >>> sitk_image = sitk.Image((224, 224), sitk.sitkVectorFloat32, 3) >>> tio.ScalarImage.from_sitk(sitk_image) ScalarImage(shape: (3, 224, 224, 1); spacing: (1.00, 1.00, 1.00); orientation: LPS+; memory: 588.0 KiB; dtype: torch.FloatTensor)
- get_bounds() Tuple[Tuple[float, float], Tuple[float, float], Tuple[float, float]][source]#
Get minimum and maximum world coordinates occupied by the image.
- get_center(lps: bool = False) Tuple[float, float, float][source]#
Get image center in RAS+ or LPS+ coordinates.
- Parameters:
lps – If
True, the coordinates will be in LPS+ orientation, i.e. the first dimension grows towards the left, etc. Otherwise, the coordinates will be in RAS+ orientation.
- property itemsize#
Element size of the data type.
- load() None[source]#
Load the image from disk.
- Returns:
Tuple containing a 4D tensor of size \((C, W, H, D)\) and a 2D \(4 \times 4\) affine matrix to convert voxel indices to world coordinates.
- save(path: str | Path, squeeze: bool | None = None) None[source]#
Save image to disk.
- Parameters:
path – String or instance of
pathlib.Path.squeeze – Whether to remove singleton dimensions before saving. If
None, the array will be squeezed if the output format is JP(E)G, PNG, BMP or TIF(F).
- set_data(tensor: Tensor | ndarray)[source]#
Store a 4D tensor in the
datakey and attribute.- Parameters:
tensor – 4D tensor with dimensions \((C, W, H, D)\).
- show(viewer_path: str | Path | None = None) None[source]#
Open the image using external software.
- Parameters:
viewer_path – Path to the application used to view the image. If
None, the value of the environment variableSITK_SHOW_COMMANDwill be used. If this variable is also not set, TorchIO will try to guess the location of ITK-SNAP and 3D Slicer.- Raises:
RuntimeError – If the viewer is not found.
- property tensor: Tensor#
Tensor data (same as
Image.data).
- to_gif(axis: int, duration: float, output_path: str | Path, loop: int = 0, rescale: bool = True, optimize: bool = True, reverse: bool = False) None[source]#
Save an animated GIF of the image.
- Parameters:
axis – Spatial axis (0, 1 or 2).
duration – Duration of the full animation in seconds.
output_path – Path to the output GIF file.
loop – Number of times the GIF should loop.
0means that it will loop forever.rescale – Use
RescaleIntensityto rescale the intensity values to \([0, 255]\).optimize – If
True, attempt to compress the palette by eliminating unused colors. This is only useful if the palette can be compressed to the next smaller power of 2 elements.reverse – Reverse the temporal order of frames.
- unload() None[source]#
Unload the image from memory.
- Raises:
RuntimeError – If the images has not been loaded yet or if no path is available.